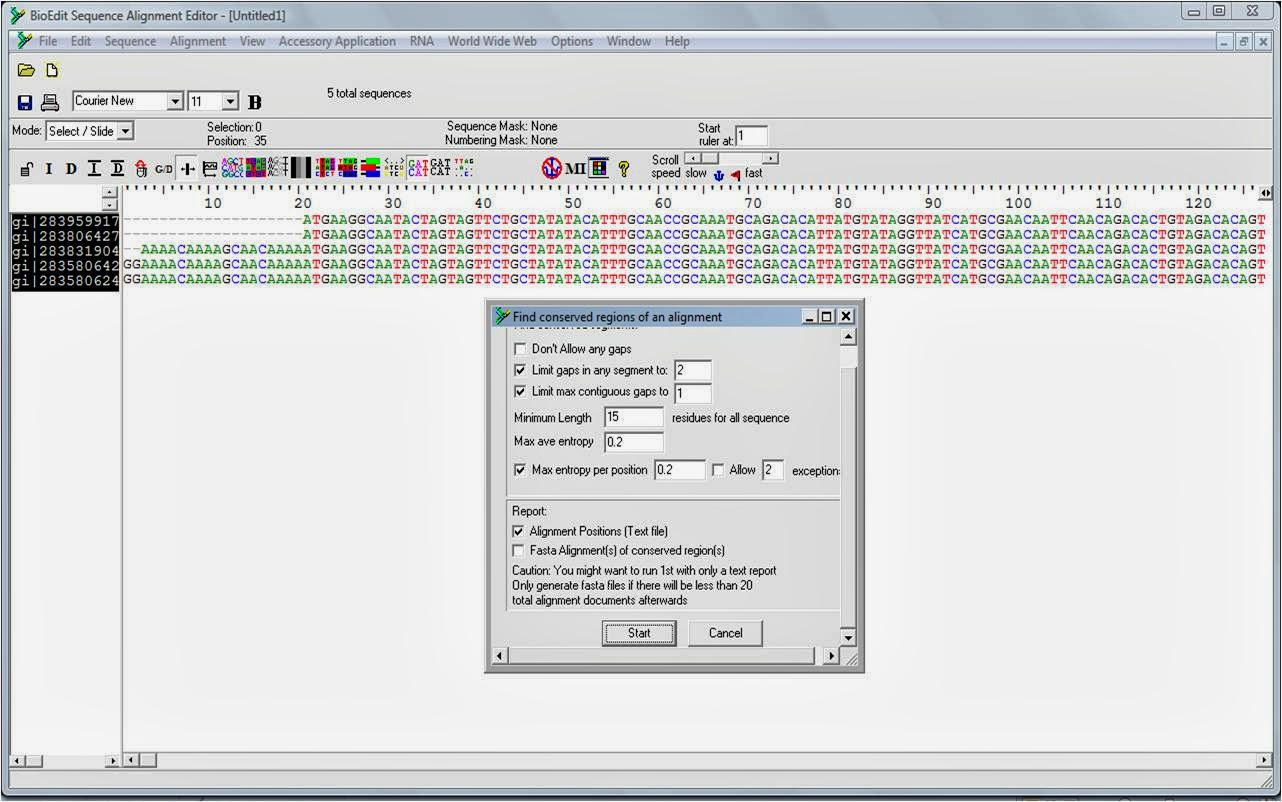
I select a point in the reverse, then select sequence to the end Edit, Select to End, control-e. The clones were sequenced using either the T7 or SP6 promoter primers that flank the multiple cloning site in this vector. This changes the way the sequences are tutoriap. Select all residues of all the sequences. I copy all the forwards to a new BioEdit file, select the sequence titles Edit, Select All Sequences, control-shift-a and copy boiedit to clipboard Edit, Copy Sequences, control-amake the new BioEdit file active and paste them in Edit, Paste Sequences, control-s. To correct the consensus sequence I copy and paste the sequences from a population or individual, group, etc. Not copied to the system clipboard.ĭrag residues with the mouse left button on. Delete and tutoria the data of highlighted sequence. Select them all control-acopy to clipboard control-cgo back to BioEdit, to paste these tutoriql over the existing ones. Once you set your preferences on one machine you can copy the bioedit.Īll of that probably sounds very confusing, once tutorail have carefully worked through it a couple of times it becomes very easy. Now I select all the forward sequences and cut them and scroll right to check for any bases changes that need to be checked. BioEdit can also edit chromatograms, biowdit I find Chromas to be nicer. To identify vector sequences, alignments will be prepared between your edited forward and reverse complement sequences and the sequence in the pstblue1vector. Adjust the size of the chromatogram trace with the Horizontal scale and Vertical scale bars to the top left of the image. I then create a second file which has only the. If the vector sequence given is the opposite strand to the forward sequence, then there should be a region of almost exact homology with the end of the reverse tuttorial. Just be sure to select to end from a different location each time to reduce the chances of pasting the wrong reverse into your consensus. Select the value you wish to change, hit the value on the keyboard and that will reset it. If I loose my sequence alignment, at least all my chromatograms with the correct edits are still there to rebuild it from. Guide to editing sequences with Chromas and BioEditĪs far as I can tell there is no difference between saving your file as a BioEdit formatted file versus as a fasta file. Indication of selected region on the aligment window not changed. If this does not occur, repeat the process with the reverse complement sequence file in a New tutoriak. Hit control-e to select to the tutorail, hit delete, move right one base then paste control-c. These should show an almost exact match to the forward or reverse sequence. Note that this is also displayed in a 5′-3′ direction, so the sequence complementary to the beginning of your original unedited forward sequence will be at the end of the reverse complement. To get the sequence of the original template strand, the Reverse Complement must be prepared.
#HOW TO EDIT SEQUENCES IN BIOEDIT PDF#
LAKSHMI NARASIMHA SWAMY SAHASRANAMAM IN TELUGU PDF Note that this works best with coding sequences without indels as every sequence is an identical length, it is all a bit trickier with different length sequences. Return to your edited forward sequence file, delete the vector sequences, and save for next week. One quirk of BioEdit is that if you double click a data file it will open in a new copy of BioEdit, not in an existing one. The chromatograms come off the machine with all bases in upper case.
#HOW TO EDIT SEQUENCES IN BIOEDIT FULL#
See sequence analysis references for full map. Eventually the forwards will start to be a poor match to the reverses. I check any unique differences by opening the chromatogram. Click on Sequence menu, Pairwise alignmentAlign two sequence allow ends to slide. On the middle toolbar 2nd in the alignment window change mode to edit, change box next to it to insert. BioEdit Tutorials – Practical Bioinformatics Save the edited file to the desktop, or preferably, your own disk or network account. The most annoying aspect is that you have tktorial manually align up each sequence and manually create a consensus sequence which commercial programs like Sequencher and Geneious are very good at. There are 4 disks containing sequence files. Įach group should choose one of the sequence files on the disk, and copy it from drive A to the desktop. MEGA also has an alignment editor, but I’ve not really used it very much. BioEdit can also edit chromatograms, but I find Chromas to be nicer. BioEdit is a mouse-driven, easy-to-use sequence alignment editor and sequence analysis program designed and written by a graduate student. This is likely to be the final release of BioEdit.
North Carolina State University, Department of Microbiology.
